How bioinformatics is tackling the life sciences data deluge: Q&A with Craig McAnulla
Craig is a bioinformatician at Eagle Genomics. In this article he discusses the scale of the data challenge in life sciences R&D and how bioinformatics technologies are evolving to enable the transformation of big data into actionable insights.
 Craig McAnulla, Bioinformatician
Craig McAnulla, Bioinformatician
"We want to help model the knowledge that is already out there so that researchers can really simply and quickly find the relevant, scientifically valid data which answers their questions."
Q: What is your role at Eagle Genomics?
I’m a bioinformatician, which means I know how to code and efficiently use computers but also have an in-depth knowledge of biology. I did a lab-based PhD in microbiology and then followed that with a couple of postdoctoral positions before I decided that I liked working with computers better than being in the lab.
During my time as a lab scientist I used the European Bioinformatics Institute’s (EMBL-EBI) data services quite a lot and I actually ended up getting a job there as a curator at the InterPro database. That’s how I started, but as time went on I got more and more involved in the computational side of things and I ended up running one of the production systems for InterPro. Eventually I was doing more programming than biology and I then joined the team at Eagle Genomics!
Q: With the production of life sciences data continuously increasing in speed and volume, what is the role of bioinformatics in tackling the data deluge?
Bioinformatics has become a confusing term in a lot of ways - it’s so broadly-defined now you could have two bioinformaticians whose jobs don’t have a single thing in common. There are bioinformaticians who analyse life sciences data, bioinformaticians who write software methodologies, those who look after databases and more.
Bioinformatics has a very important role in tackling the huge volumes of life sciences data out there. It’s a difficult challenge which is going to require more technical expertise from computer scientists due to the level of infrastructure needed to tackle data at this scale. But it also requires domain experts in biology; bioinformaticians are essential for marrying the technical side of life sciences data management with the end users, the lab scientists.
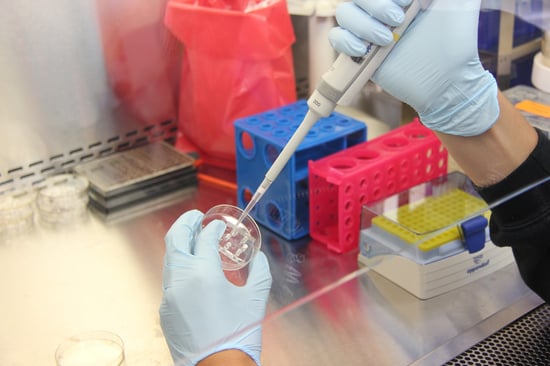 Bioinformaticians are key to managing the technical aspect of life sciences data management.
Bioinformaticians are key to managing the technical aspect of life sciences data management.
Q: What are the specific challenges of data analysis in a life sciences context?
The first challenge is just finding the data! That’s a real issue. There’s loads of data out there, but how can scientists find the data they need? It could be publically available but even if it’s in-house, within a company’s private data, researchers still need to be able to locate the data and understand enough about its context to be able to access and analyse it meaningfully.
Another issue is the sheer volume of life sciences data. How can researchers analyse such huge amounts of data within a reasonable timeframe? If a researcher has to perform their analyses in the cloud they have to pay for that service, so they need to choose analyses which are fast enough to be cost effective but also of good enough quality to give meaningful results! So sometimes there is a trade-off to be made between the optimal analysis, which would give the best possible results, versus what’s actually practical to run on the data volume.
Additionally, after a pipeline analysis is performed the results of that pipeline might actually be bigger in volume than the original dataset! Researchers can end up with a large amount of data very, very quickly which is why they end up using systems like the cloud because this makes it much easier to scale storage as required.
Q: Why are effective data analysis methods so important for R&D?
Our understanding evolves all the time; the way a metagenomic experiment is analysed now is not the same as it was ten years ago, which is a good thing because methods have become faster and more effective.
When researchers use a bioinformatics pipeline there are technical elements to that pipeline which aren’t really doing any biology but are essential for generating meaningful results. For example, quality trimming and filtering of data isn’t really biological or an end-result researchers are interested in, however it is cleaning up the data, which is absolutely fundamental to producing meaningful analysis.
 A researcher may want to investigate how a specific condition affects the skin microbiome.
A researcher may want to investigate how a specific condition affects the skin microbiome.
Clean data then enables the scientific questions to come into play. For example, a researcher may want to find out if sample A is different to sample B, or if a specific treatment affects the skin microbiome significantly. High quality technical elements within a pipeline, such as sequence denoising and data filtering, enable researchers to capture reliable scientific results much more quickly and accurately.
Q: Research data relevant to a specific area of interest often exists outside of an organisation as well as internally. How can the challenge of disparate datasets be overcome to provide the most informed view of an area of research?
People are working on this, initiatives like the FAIR data sharing principles are going to be really important for ensuring that data is standardised and unifiable. Data curation and the use of standardised ontologies across databases are fundamental to enabling the conversation between the data and the scientist.
Another challenge is that experiments these days involve multi-omics data. These different data types, although they are all from a single experiment, might be stored in different databases. How is a user then easily able to access all those different data types? At the moment this still presents a significant obstacle, so I would encourage anyone to support initiatives like ELIXIR and FAIR and to lobby on this issue for more funding to try to enhance the accessibility of data.
Once standards have been realised we can apply much more effective valuation and contextulisation in order to unify multi-omics datasets.
Q: How can Eagle’s e[datascientist] help researchers channel the data deluge into actionable insights?
A user could come to the system and ask the question ‘Which skin microbiome is associated with dandruff?’ in order to help them develop a treatment for that condition.
In that situation they would want to find out which scalp microbiome sets are available and if there are any pre-existing data in this area which they can look at. Currently this is difficult for researchers. The Eagle Genomics platform makes this easier by integrating in-house datasets with other relevant data identified in open source databases.
The platform will also provide the capability to identify any existing analysis results which other researchers have produced using this data. Within the platform a user will then be able to run analysis pipelines on their chosen data and follow that up with statistical analyses.![e[datascientist]](https://www.eaglegenomics.com/hs-fs/hubfs/Imported_Blog_Media/e%5Bdatascientist%5D-2.png?width=600&name=e%5Bdatascientist%5D-2.png)
Visulisation of the e[datascientist] from big data to insight
A user will be able to go end-to-end, from collating the starting data to discovering biological insights. We want to help model the knowledge that is already out there so that researchers, including non-data-science-experts, can really simply and quickly find the relevant, scientifically valid data which answers their questions.
Q: What are your hopes for the advancement of data analysis within the life sciences?
It’s difficult to say. I’ve already seen so many things happen in the field which I didn’t expect. I remember when the E.coli genome was sequenced and that was a massive deal, every time a new genome was sequenced it was very exciting. Nowadays it's just trivial!
When I did my PhD I worked on a bacterium that hadn’t been sequenced. If we’d had the genome sequence we could have probably done my whole PhD in a couple of months!
It can still be a struggle for scientists to find the information that they really need, or to even know that it exists in the first place. That's something Eagle Genomics is committed to resolving with the e[datascientist] platform.
The more we can do to make a lab scientist's life simpler and give them what they need to do their job, the better. If you can speed somebody’s work up from two weeks to five minutes, that’s more experimental work they can be doing in order to develop novel treatments and solutions.
![Learn more about e[datascientist]](https://no-cache.hubspot.com/cta/default/2869527/95a273e6-3721-4619-96f8-c62aaf8fade1.png)