Are household products contributing to antibiotic resistance?
Nowadays, antiseptics and disinfectants are widely used by the general public to clean various contact surfaces, including taps, doorhandles, kitchen work tops and bathroom sinks and toilets. These reagents demonstrate broad-spectrum activity, and claim to kill 99.9% of bacteria and viruses (e.g. E. coli, MRSA).
However, some studies suggest that more extensive and longer term use of certain disinfectants (e.g. triclosan) might select for micro-organisms that are antibiotic resistant (i.e. micro-organisms that acquired the ability to resist antibiotics that would normally kill them or stop their growth). This is an urgent threat as these ‘superbugs’ can cause life threatening illnesses due to infection, and since they are antibiotic resistant, they are very difficult to treat.
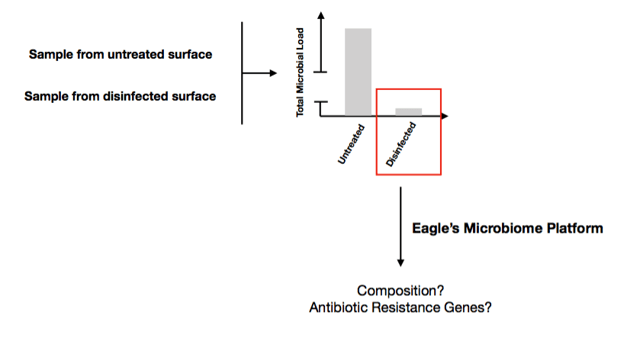
Therefore, it is important that, although 99.9% of the “biological sample” is killed upon disinfecting, the remaining 0.1% are monitored for the presence of antibiotic-resistance genes.
Eagle’s microbiome platform now allows for fast and accurate detection of antibiotic-resistance genes within your metagenome sample by integrating available deep-learning approaches into our microbiome analysis workflow. This means that besides determining the composition of your sample, this workflow also identifies genes encoding known antibiotic resistance mechanisms.
For more on our microbiome platform, join our upcoming webinar 'Expert Panel on Industrialising Microbiomics', with guest panelist, Dr Barry Murphy, Microbiomics Science Lead at Unilever.
